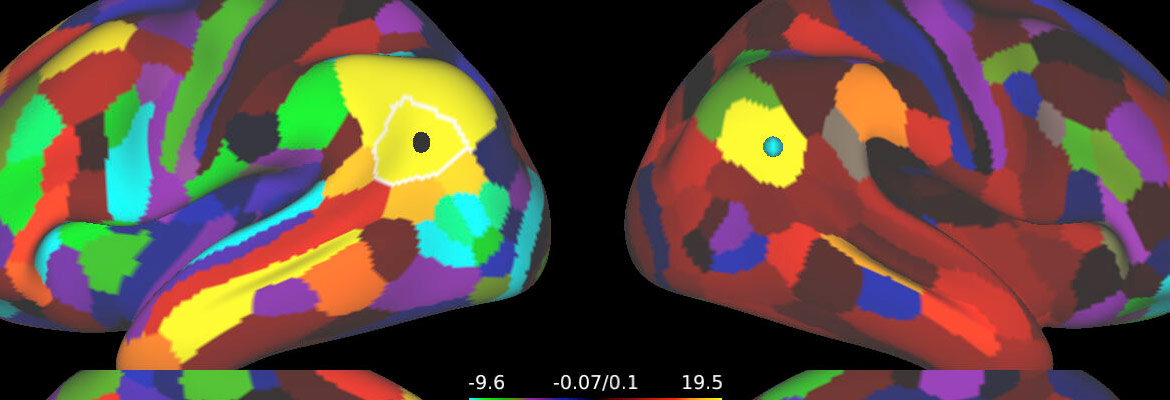
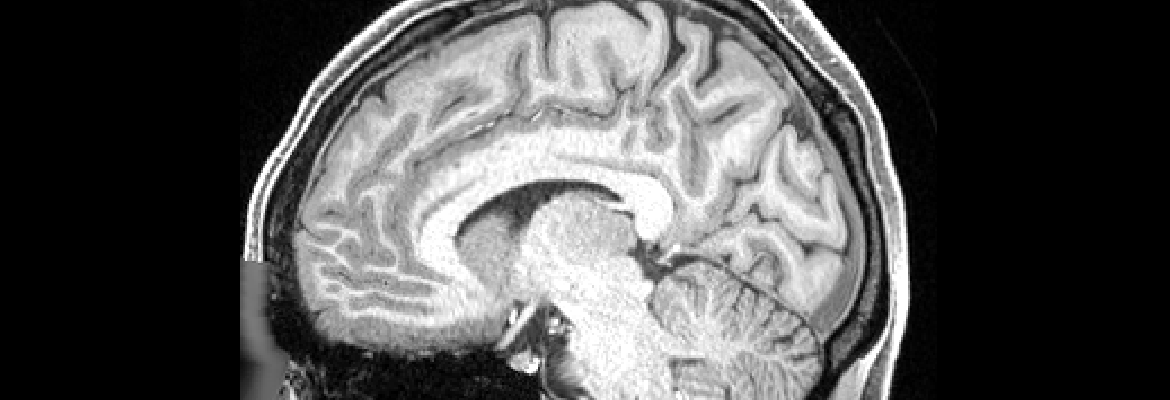
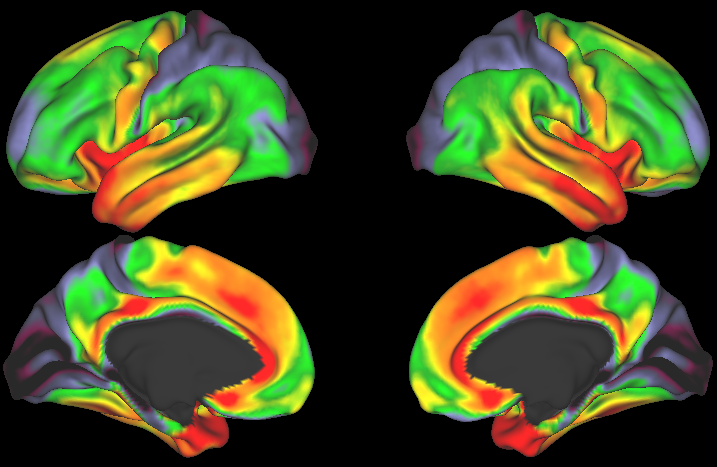
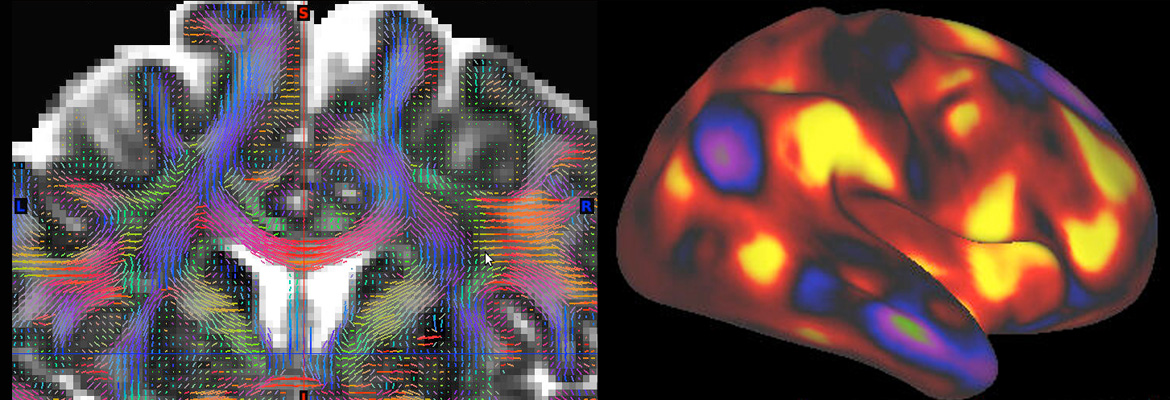
The Human Connectome Project (HCP) has tackled one of the great scientific challenges of the 21st century: mapping the human brain, aiming to connect its structure to function and behavior.
What is the Connectome Coordination Facility?
The Connectome Coordination Facility (CCF) houses and distributes public research data for a series of studies that focus on the connections within the human brain. These are known as Human Connectome Projects.
The CCF currently supports 20 human connectome studies. Scroll down to learn more.
Lifespan HCP
HCP Lifespan Projects are acquiring and sharing multimodal imaging data acquired across the lifespan, in four age groups (prenatal, 0-5, 6-21, and 36-100+).
Connectomes Related To Human Disease
HCP Disease studies apply HCP-style data collection methods to subject cohorts at risk for, or suffering from, disorders affecting the brain.
HCP Software
We have released and maintain a set of open-source Connectome software that supports browsing, download, exploration, visualization and analysis of HCP data.
Healthy Adult Connectomes
The Human Connectome Project (HCP) has tackled one of the great scientific challenges of the 21st century: mapping the human brain, aiming to connect its structure to function and behavior.
HCP Young Adult
PI: Kamil Ugurbil, David Van Essen1200 Subjects, Age 22-35
Washington U. in Saint Louis, U. of Minnesota, U. of Oxford, Saint Louis U., Indiana U., U. d’Annunzio, Ernst Strungmann Institute, Warwick U., Radboud U. Nijmegen, U. of California at Berkeley
Current Data Releases (All Data Releases)
Lifespan Connectome Data
HCP Lifespan Projects are acquiring and sharing multimodal imaging data acquired across the lifespan, in four age groups (prenatal, 0-5, 6-21, and 36-100+). The scanning protocols are similar to those for the WU-Minn Young Adult HCP, except shorter in duration.
HCP Aging
PI: Beau Ances, Susan Bookheimer, Randy Buckner, David Salat, Stephen Smith, Melissa Terpstra, Kamil Ugurbil, David Van Essen, Roger Woods1200 Subjects, Age 36-100+
Washington University, University of Minnesota, Massachusetts General Hospital, Harvard University, University of California Los Angeles, Oxford University
Current Data Releases (All Data Releases)
HCP Development
PI: Deanna Barch, Susan Bookheimer, Randy Buckner, Mirella Dapretto, Stephen Smith, Leah Somerville, Kathleen Thomas, David Van Essen, Essa Yacoub1350 Subjects, Age 5-21
Washington University, University of Minnesota, University of California at Los Angeles, Harvard University, Oxford University
Current Data Releases (All Data Releases)
Lifespan Baby Connectome Project
PI: Jed Elison, Weili Lin500 Subjects, Age 0-5
University of North Carolina, University of Minnesota
Lifespan Developing Human Connectome Project
PI: David Edwards, Jo Hajnal, Daniel Rueckert, Stephen Smith1500 Subjects, Age 20-44 weeks post-conception
Connectomes Related To Human Disease
HCP Disease studies apply HCP-style data collection protocols toward subject cohorts at risk for, or suffering from, diseases or disorders affecting the brain, with a goal of providing comparable data to healthy HCP subjects across the lifespan.
Alzheimer's Disease Connectome Project
PI: Barbara Bendlin, Shi-Jiang Li300 Subjects, Age 55-90
Medical College of Wisconsin, University of Wisconsin
Amish Connectome Project
PI: Elliot Hong, Peter Kochunov450 Subjects, Age 18-85
University of Maryland
BANDA: Connectomes Related to Anxiety & Depression
PI: John Gabrieli, Susan Whitfield-Gabrieli225 Subjects, Age 14-17
Massachusetts Institute of Technology, Massachusetts General Hospital, McLean Hospital, Boston University
Current Data Releases (All Data Releases)
Changes in Visual Cortical Connectivity Following Central Visual Field Loss
PI: Kristina Visscher100 Subjects, Age 18-89
University of Alabama
Connectomic Imaging in Familial & Sporadic Frontotemporal Degeneration
PI: Murray Grossman, Corey McMillan200 Subjects, Age 18-60
University of Pennsylvania, Mayo Clinic, Massachusetts General Hospital, Northwestern University, University of California San Francisco
Connectomics in Brain Aging and Dementia
PI: James Becker400 Subjects, Age 50-89
University of Pittsburgh
Dimensional Connectomics of Anxious Misery
PI: Yvette Sheline250 Subjects, Age 18-45
University of Pennsylvania
Current Data Releases (All Data Releases)
Epilepsy Connectome Project
PI: Jeff Binder, Beth Meyerand340 Subjects, Age 18-50
Medical College of Wisconsin, University of Wisconsin
Human Connectome Project for Early Psychosis
PI: Alan Breier, Martha Shenton400 Subjects, Age 16-35
Boston, MA: Brigham and Women's Hospital, Beth Israel Deaconess-Massachusetts Mental Health Center, McLean Hospital, Massachusetts General Hospital;
Indianapolis, IN: Indiana University
Current Data Releases (All Data Releases)
Human Connectomes for Low Vision, Blindness, and Sight Restoration
PI: Geoffrey Aguirre, Vivek Patel, Yonggang Shi260 Subjects, Age 20-80
University of Southern California, University of Pennsylvania
Mapping Connectomes for Disordered Emotional States (HCP-DES)
PI: Leanne Williams300 Subjects, Age 18-35
Stanford University
Neural Disconnection & Errant Visual Perception in Psychotic Psychopathology
PI: Scott Sponheim300 Subjects, Age 18-59
University of Minnesota
Perturbation of the Treatment of Resistant Depression Connectome by Fast-Acting Therapies (PDC)
PI: Randall Espinoza, Katherine Narr, Danny Wang240 Subjects, Age 20-64
University of California, Los Angeles
Current Data Releases (All Data Releases)
The Structural & Functional Connectome across AD Subtypes
PI: John Ringman260 Subjects, Age 20-80
University of Southern California
Related Connectome Studies
These studies are not explicitly chartered under the Lifespan HCP or Disease HCP grants, but are studying brain function using the rigor and methods of the Human Connectome Project.
Dual Mechanisms of Cognitive Control
PI: Todd Braver300 Subjects, Age 18-40
Washington University School of Medicine
Connectome Software Releases
Connectome software has been developed that fully supports browsing, downloading, exploring and analyzing of HCP data. Connectome software will be downloadable as individual components with full documentation on installing and using the tools.
Connectome Workbench
v 2.0.0
Screenshots
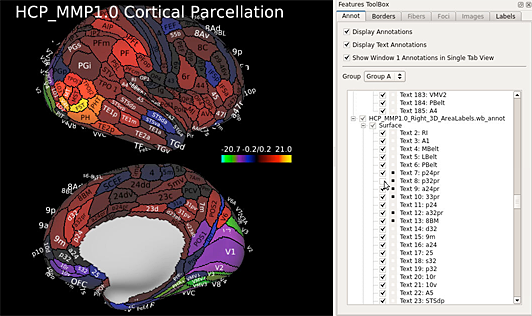
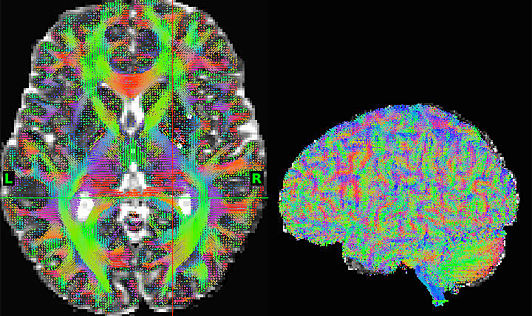
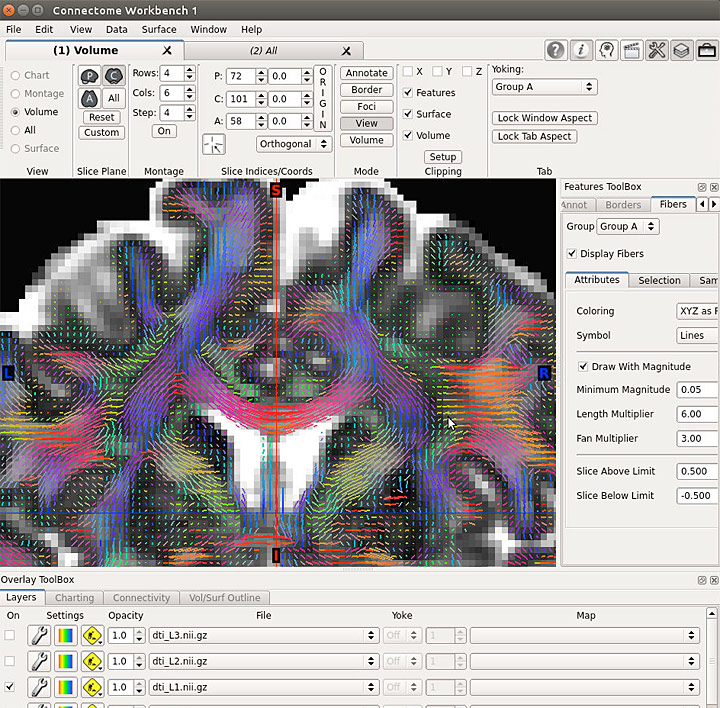
Connectome MR Pipeline
v 4.7.0
The HCP Pipelines product is a set of tools (primarily, but not exclusively, shell scripts) for processing MRI images for the Human Connectome Project. Among other things, these tools implement the Minimal Preprocessing Pipeline (MPP) described in Glasser et al. 2013.
HCP MEG Pipelines
v 3.0
The analysis of MEG data in the Human Connectome Project is performed using FieldTrip, a MATLAB toolbox for MEG and EEG analysis, in combination with additional analysis scripts and functions that have specifically been written for the HCP.