Using the Connectome
Connectome Software
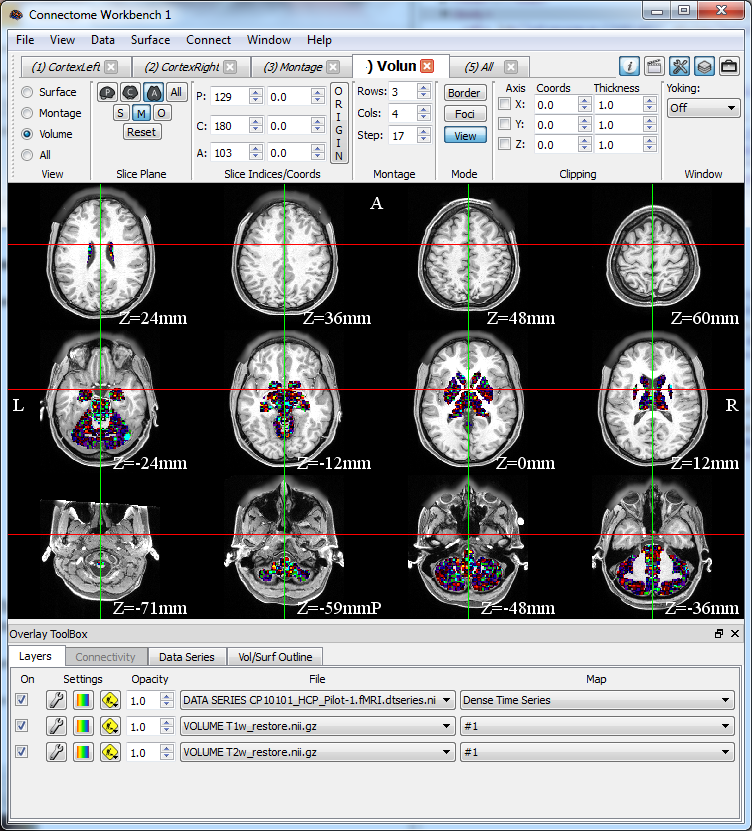
The Human Connectome Project consortium has developed software tools as a part of our HCP Informatics Infrastructure to support browsing, download, exploration and analysis of connectome data. Additionally, the CCF has partnered with the NIH Data Archive to widely distribute CCF study data.
- All data from CCF studies, including the original Human Connectome Project young adult dataset, will be available for download from the NIH Data Archive.
- All connectome data can be imported seamlessly into Connectome Workbench, which is used to visualize and analyze connectivity data in individuals or population-averaged brain atlases, on surfaces and in volumes.
HCP Pipelines
HCP pipelines for both MR and MEG processing have been publicly released.
- MR Pipelines: Pipeline scripts implement the Minimal Preprocessing Pipeline (MPP) described in Glasser et al. 2013 (including structural processing by FreeSurfer), single and multi-run ICA-FIX cleaning of fMRI data, task analysis using FEAT, diffusion analysis using bedpostX and 7T data processing.
- MegConnectome: The analysis of MEG data in the Human Connectome Project is performed using FieldTrip, a MATLAB toolbox for MEG and EEG analysis, in combination with additional analysis scripts and functions that have specifically been written for the HCP.
HCP Analysis Tools
The Connectome toolbox also includes analysis tools such as:
- FSL is a comprehensive (structural, functional and diffusion) neuroimaging software platform. FSL tools will be used in a variety of contexts in the HCP, including preprocessing, diffusion analysis and tractography, R-fMRI ICA, and parcellation.
- FreeSurfer is used for structural data analysis, including generation of cortical surfaces and subcortical segmentation.
- Brain Connectivity Toolbox (BCT) provides an access to a large selection of complex network measures in Matlab. Such measures aim to characterize brain connectivity by neurobiologically meaningful statistics, and are increasingly used in the description of structural and functional connectivity datasets
- Fieldtrip is an open source Matlab toolbox for MEG, EEG and LFP analysis. It includes algorithms for simple and advanced analysis of electrophysiological data, such as time-locked averaging, time-frequency analysis, source reconstruction using dipoles, distributed sources and beamformers, channel and source-level synchronization estimates and non-parametric statistical testing.