- >>
- Healthy Adult Studies
- HCP Young Adult
- Project Protocols Detail
Components of the Human Connectome Project - Multi-Modal Data Integration
The Human Connectome Project involves the acquisition of imaging data sets from multiple modalities (HARDI, R-fMRI, T-fMRI, MEG) across a large number of individuals. Integrating these multimodal data sets is critical for achieving several key scientific goals: a) the reliable and robust parcellation of the brain into a set of distinct regions; b) the registration and comparison of structural and functional connectivity across imaging modalities, as well as across individual subjects.
Brain parcellation is made difficult because of the large number of regions and pathways, the high level of individual variability in brain anatomy, the complex relationship between regional boundaries and interregional connectivity, and limits on the spatial resolution of all imaging methods. New techniques are needed to overcome these challenges, to achieve reliable parcellation of the whole brain, a key prerequisite for subsequent network modeling and analysis. These approaches are carried out in parallel on HARDI, R-fMRI and T-fMRI data sets. Using structural connectivity data acquired with HARDI, we apply connectivity-based parcellation techniques, implementing the idea that neural elements forming a coherent brain region will share similar connectivity patterns to other regions in the brain. Using R-fMRI data, we will extract boundaries where functional connectivity patterns exhibit significant discontinuities, creating gradient maps that define regions with internally coherent functional connectivity. Both approaches use connectivity (structural and functional) to achieve whole brain parcellation and utilize a variety of clustering strategies and algorithms. Another promising approach to extract separate spatial components corresponding to single regions or sets of regions from imaging data, called independent component analysis (ICA), is being optimized for use in single-subject and multi-subject parcellation.
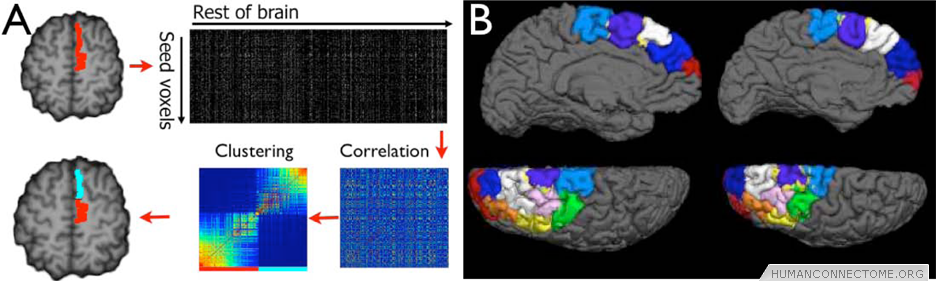
Figure 1: (A) tractography parcellation. Connectivity distributions are calculated from seed voxels. Correlations are computed resulting in a similarity matrix between seed voxels. Clusters are identified in this similarity matrix and mapped back into the brain. (B) This approach has been applied to large tracts of cortex – Here showing remarkable consistency in parcellating the dorsal prefrontal cortex in two subjects.
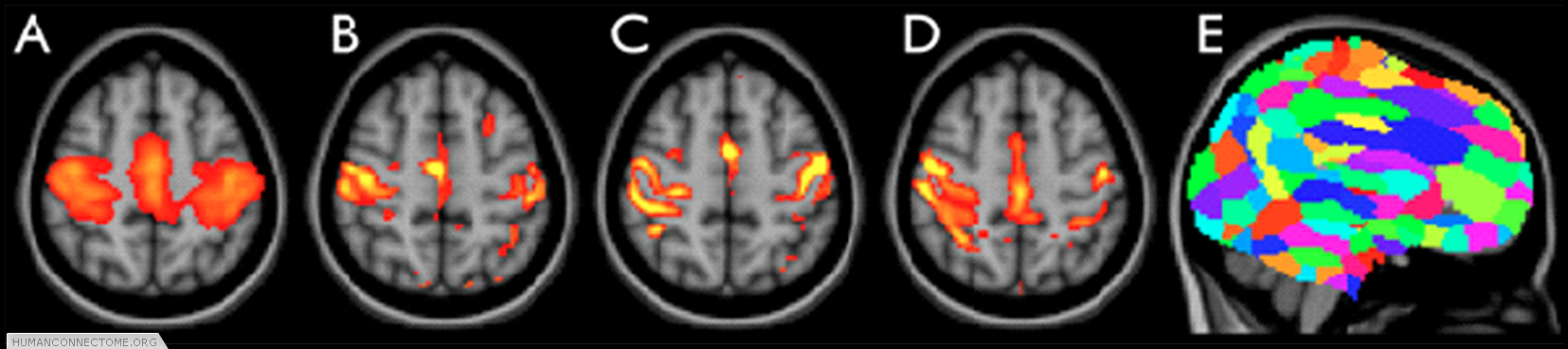
Figure 2: ICA-based parcellation of 3T resting-FMRI data. (A) group-mean parcellation of a primary sensori-motor region (one “resting state network”). (B-D) The corresponding map from 3 different subjects, achieved by applying “dual regression” of the group map into each individual data set. (E) Whole-brain parcellation resulting from applying ICA to the group data and combining across all ICA components.
These different parcellation strategies are being evaluated in several ways: by testing for consistency across repeated scans and across subjects, by comparing across 3T and 7T data sets, by checking for alignment of partitions obtained with HARDI, R-fMRI, and T-fMRI, by applying network metrics capturing the modularity of brain connectivity, and by comparison of population parcellations to registered cytoarchitectonic maps. Brain parcellations, for example those derived from R-fMRI, are also being compared with patterns of T-fMRI activations, thus establishing that the identified regional partitions correspond to sets of brain regions that are functionally engaged across different behavioral and cognitive tasks.
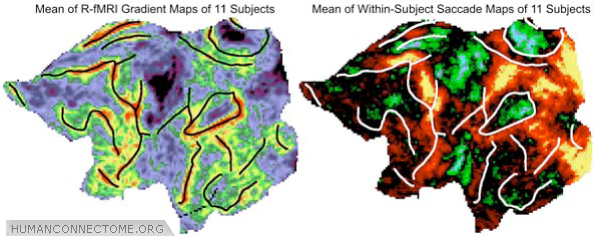
Figure 3: Comparison of major R-fMRI gradient boundaries (left) to T-fMRI activation patterns obtained during an eye movement task (right), displayed here in a population average (n = 11).
The comparison of parcellation and connectivity patterns obtained from different imaging modalities can aid in defining more robust regional partitions. Several studies have documented that structural and functional connectivity in the human brain is statistically related, likely reflective of the fact that anatomical pathways shape and constrain dynamic interactions among brain regions. These relationships can be exploited to more reliably identify functional brain regions in single individuals. For example, the integration of gradient maps corresponding to putative boundaries in structural and functional connectivity is expected to yield more reliable and consistent whole-brain parcellation. ICA approaches can also be applied in a multimodal context, by comparing ICA-based partitions obtained from HARDI and R-fMRI data, before developing methods for joint decomposition.
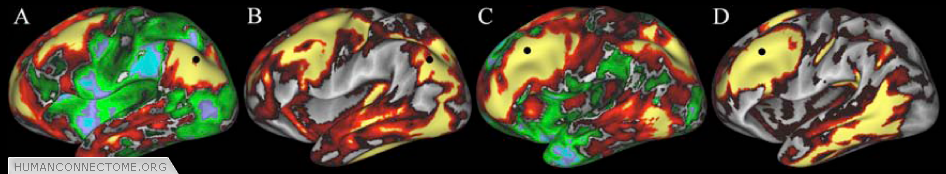
Figure 4: Similarities between R-fMRI (A,C) and diffusion imaging (B,D) connectivity maps. Connection patterns for each modality are displayed concurrently on paired surfaces using a ‘point-and-click’ option in Caret. Seed point for functional connectivity and tractography (black circle) is the same in A and B, and in C and D.
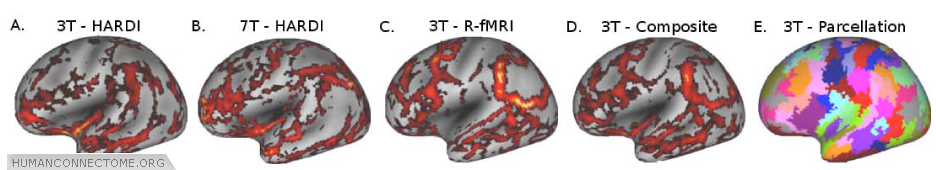
Figure 5: A and B. Structural connectivity gradient maps (thresholded) from 1 subject scanned at 3T and another scanned at 7T (Correlation coefficient = 0.26, p<10-4). C. Subject A’s R-fMRI connectivity gradient maps. D. Gradient map generated by combining structural and R-fMRI connectivity information from Subject A. E. Cortical partitioning based on K-means clustering using local minima from the composite gradient shown in D.
A variety of techniques are being refined and further optimized to achieve accurate within-subject alignment of data sets of multiple modalities, as well as precise inter-subject registration. Our expertise in image distortion correction and cross-modal registration allow us to achieve reliable and precise alignment of structural and functional imaging data in single subjects. Volume, surface and connectivity-based approaches to cross-subject registration are being concurrently explored and compared.
Further Reading:
Beckmann, C.F. and S.M. Smith (2004) Probabilistic independent component analysis for functional magnetic resonance imaging. IEEE Trans Med Imaging. 23: 137-52.
- Cohen, A.L., D.A. Fair, N.U. Dosenbach, F.M. Miezin, D. Dierker, D.C. Van Essen, B.L. Schlaggar and S.E. Petersen (2008) Defining functional areas in individual human brains using resting functional connectivity MRI. Neuroimage. 41: 45-57.
Johansen-Berg, H., T.E. Behrens, M.D. Robson, I. Drobnjak, M.F. Rushworth, J.M. Brady, S.M. Smith, D.J. Higham and P.M. Matthews (2004) Changes in connectivity profiles define functionally distinct regions in human medial frontal cortex. Proc Natl Acad Sci U S A. 101: 13335-40.
Smith, S.M., P.T. Fox, K.L. Miller, D.C. Glahn, P.M. Fox, C.E. Mackay, N. Filippini, K.E. Watkins, R. Toro, A.R. Laird, and C.F. Beckmann (2009) Correspondence of the brain's functional architecture during activation and rest. Proc Natl Acad Sci U S A. 106: 13040-5.